|
It is often useful to organise groups of components within a model into a hierarchical structure. This structure might reflect the logical organisation of components within the group or their physical configuration. CellML provides a single mechanism for the specification of both of these forms of hierarchy. This mechanism is based on a grouping scheme that allows model authors to create numerous hierarchical structures over a single network of components. The parent-child relationships in one hierarchical grouping need not necessarily be consistent with those specified in another grouping, a situation that could not be supported by nesting of component definitions.
It is anticipated that models will typically be defined as a network, with hierarchical relationships defined between groups of components at different places within the model. CellML processing software is free to treat these structures as discontinuous. Alternatively, it may combine structures that represent the same relationship into a single hierarchy by assuming that the root nodes of any hierarchical arrangements of components are all children of a single imaginary component. This imaginary component is not explicitly defined within the CellML document and has no properties.
The definition of a logical hierarchy of components in a network is known as "encapsulation". Encapsulation allows the modeller to hide a group of components from the rest of the model by using a single component as an interface to the hidden subnetwork. The parent component hides the details of one or more child components from the rest of the model. Encapsulation provides a powerful mechanism for simplifying the structure of the model by preventing connections between specified sets of components. Components in the main network may not connect to the child components in the subnetwork — all variables must be mapped through the parent interface component. Components in the subnetwork may only be connected to the interface component and to other components in the same subnetwork, which may include further levels of encapsulation. Therefore, a modeller wishing to re-use an encapsulated subnetwork may treat the subnetwork as a "black box", and deal exclusively with the interface presented by the encapsulating component.
The definition of physical hierarchies within a model is known as "containment" in CellML. A model author can specify that one or more child components are physically inside of a parent component without describing the geometric aspects of the relationship in detail. This information would typically be used by CellML processing software to provide simple renderings of a model.
Model authors are also free to extend the grouping scheme with user-defined types of relationships between components. However, CellML processing software is only expected to recognise encapsulation and containment relationships.
Groups do not add any additional mathematical information to the model. Model authors may not define their own grouping relationships that are intended to imply mathematical information.
Logical and physical hierarchies are both declared using the <group> element. This element must be a child of a <model> element. Each <group> element contains one or more <relationship_ref> elements, each of which defines a relationship attribute, the value of which references the type of relationship represented by the group. CellML processing software is expected to recognise two types of relationship: encapsulation and containment, which are indicated by relationship attribute values of "is_encapsulated_by" and "is_contained_in", respectively.
The <group> element also contains two or more <component_ref> elements, each of which defines two attributes. The component attribute references a component within the current model. The role attribute indicates whether the component is the dominant component in the hierarchy. A component referenced by a <component_ref> element with a role attribute value of "major" is the dominant component. This component is the encapsulating component in a logical encapsulation hierarchy or the containing component in a geometric containment hierarchy. A <group> element contains one or more <component_ref> elements with a role attribute value of "minor". The components referenced by these elements are the encapsulated components in a logical encapsulation hierarchy or the contained components in a geometric containment hierarchy. A <group> element that defines a logical encapsulation or geometric containment relationship must reference exactly one major component and at least one minor component. A group element that defines a user-defined type of relationship may have any number of minor and major components.
A single <group> element may be used to define multiple relationships between components. For instance, encapsulation and geometric relationships may be defined within the same <group> element and thus share the same hierarchy. This is done by including more than one <relationship_ref> element in the <group> element. Each <relationship_ref> element must define a relationship attribute, which may be in the CellML namespace or in an extension namespace. The value of the relationship attribute names the type of relationship referenced by the <relationship_ref> element. If the relationship attribute is in the CellML namespace, its value must be either "is_encapsulated_by" or "is_contained_in". A <relationship_ref> element may also define a name attribute. The value of the name attribute on <relationship_ref> elements can be used to combine several <group> elements into a single hierarchical structure (see Section 6.2.4 for more information on this).
Geometric containment relationship information is formally independent of logical encapsulation information, but CellML processing software is free to check for inconsistencies between the two relationships — it would generally not be useful for an encapsulating component to be physically inside one of its encapsulated child components.
All children of a given major component in a single hierarchy must appear within a single <group> element. This simplifies the construction and validation of hierarchies from <group> elements. For instance, the requirement that a component may only have a single parent in any given hierarchy would be difficult to enforce if minor components could be scattered across several <group> elements.
Encapsulation allows the modeller to split a model into layers of complexity. A single component can be used to encapsulate a complex partial model, and thereby provide a unified interface for all information passing between that subnetwork and the rest of the model.
A model may only define a single encapsulation hierarchy, which may be continuous or discontinuous. Each component in the hierarchy may have at most one parent component. If the hierarchy is continuous, the parent component will always be another component defined within the current model. If the hierarchy is discontinuous, it may be convenient to assume that any unencapsulated components are children of a single imaginary component. This imaginary component makes it easier to check that the hierarchy has no circular relationships between components.
The components in a model can be divided into four sets with respect to any given component (the current component). The set of all components immediately encapsulated by the current component is the encapsulated set. The parent component is the component that encapsulates the current component. Other components encapsulated by the same parent make up the sibling set. All other components, which are not available to make connections with the current component, make up the hidden set. If the current component is not encapsulated, then it has no parent and the sibling set consists of all other unencapsulated components in the model.
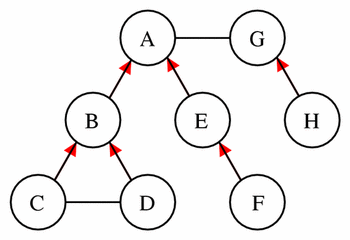
These sets are best demonstrated by example. Given the network shown in Figure 10, Table 4 lists the parent components and the components in the encapsulated, sibling, and hidden sets for a selected set of components picked as the current component.
Figure 10
This simple model provides the basis for the demonstration of the concepts of encapsulated sets, parents, sibling sets, and hidden sets, as described in the text. The model consists of eight components each represented by a circle. The lines between the components represent connections, and a red arrowhead on one of these lines indicates that the component at the tail of the arrow is encapsulated by the component at the head of the arrow.
| Current Component | Encapsulated Set | Parent | Sibling Set | Hidden Set |
|---|
| A | B, E | imaginary | G | C, D, F, H | | B | C, D | A | E | F, G, H | | C | none | B | D | A, E, F, G, H | | E | F | A | B | C, D, G, H | | G | H | imaginary | A | B, C, D, E, F |
Table 4
This table lists the parent components, and the components in the encapsulated, sibling, and hidden sets for a selected few components from the example model in Figure 10. Components A and G do not have a real parent component, but may have an imaginary parent component that enables the formation of a single encapsulation hierarchy.
Every variable must define its availability for use in other components. This is done with the public_interface and private_interface attributes on the <variable> element. The interface exposed to the parent component and components in the sibling set is defined by the public_interface attribute. The private_interface attribute defines the interface exposed to components in the encapsulated set. Each interface has three possible values: "in", "out", and "none", where "none" indicates the absence of an interface. The separation of interfaces allows the modeller to incrementally add complexity to a encapsulated network without changing the interface presented to the rest of the network by the encapsulating component.
The mappings that are allowed between variables declared in each component are controlled by the public and private interfaces of each variable and the prohibition on connecting an encapsulated component to components other than its parent component, members of its sibling set, and any components it in turn encapsulates. Variables with a public_interface attribute value of "in" must be mapped to a single variable in the sibling set with a public_interface attribute value of "out" or to a single variable in the parent of the current component with a private_interface attribute value of "out". Similarly, variables with a public_interface value of "out" may be mapped to variables in components in the sibling set with a public_interface attribute value of "in" or to variables in the parent component with a private_interface value of "in". Note that defining a public_interface attribute value of "out" on a variable makes it legal to map the variable to other variables, but does not require that such a mapping occur. If a variable has a public_interface attribute value of "none", it cannot be mapped to variables in the parent component or to variables in components in the sibling set.
Variables with a private_interface attribute value of "in" must be mapped to a single variable from a single component in the encapsulated set with a public_interface attribute value of "out". Variables with a private_interface attribute value of "out" may be mapped to any variables from components in the encapsulated set with a public_interface attribute value of "in". If a variable has a private_interface attribute value of "none", it is neither input from or exposed to the components in the encapsulated set.
If both the public_interface and private_interface attributes of a variable have a value of "none", the variable can only be used in the current component and is invisible to all other components in the model. In order to determine which variables may be modified in the current component, we must determine if either the public_interface attribute or the private_interface attribute has a value of "in". If so, the variable is declared elsewhere and its value may not be mathematically modified in the current component. If not, the variable belongs to the current component.
The two interface attributes of a variable are completely independent with one exception: it is invalid for a variable to have both public_interface and private_interface attributes with value "in". An interface with value "in" reflects an unmet need in the current component that must be satisfied — this need can be met in either the public or private interface, but not both.
The is_contained_in relationship allows the modeller to specify that a particular component is physically inside another. This might be used by software for the rendering of a model. Containment relationships can be specified either in combination with or independent of encapsulation relationships. Containment relationships do not restrict any aspect of model definition or behaviour.
CellML allows the definition of multiple containment hierarchies over the same network model. This functionality allows the modeller to define several different ways of organising the same model, each of which might highlight a different aspect of the model's physical structure. This functionality has been included in CellML for extended compatibility with AnatML, an XML-based language for describing anatomical structures.
A containment hierarchy is created when several <group> elements contain <relationship_ref> elements with a relationship attribute value of "is_contained_in" and the same name attribute value. Any <group> elements that contain <relationship_ref> elements with a relationship attribute value of "is_contained_in" and that do not define a name attribute are also considered to form a single grouping hierarchy.
As was the case for encapsulation grouping, a containment hierarchy may be continuous or discontinuous. Each component in the hierarchy may have at most one parent component. If the hierarchy is continuous, the parent component will always be another component defined within the current model. If the hierarchy is discontinuous, it may be convenient to assume that any components not already contained within other components are children of a single imaginary component. This imaginary component makes it easier to ensure that the hierarchy has no circular relationships between components.
Modellers are free to use the grouping syntax of CellML to organise model components in ways not described in the CellML specification. To do this, the model author defines a new relationship type, the name of which is used as the value of the relationship attribute on the <relationship_ref> element. The relationship attribute must be placed in an extension namespace, because future versions of the CellML specification may define additional relationship types, the names of which could otherwise conflict with user-defined relationship types. If a modeller uses a non-standard value for the relationship attribute, the value used should indicate the relationship between minor and major components. A <group> element that defines a user-defined type of group is free to contain only minor components. For example, a modeller may define a grouping class called "is_next_to", used to tell a processor that one minor component is physically adjacent to another.
Modellers are free to use the name attribute on the <relationship_ref> element to specify multiple hierarchies for user-defined relationship types, as is possible for the containment relationship.
This specification does not provide a mechanism by which modellers may specify the meaning of a user-defined type of relationship. This definition must be provided by the processing software declaring the new relationship type.
Figure 11 demonstrates the use of the <group> element to define an encapsulation relationship. This example is taken from the two reaction pathway with encapsulation example from the examples section of the CellML website. It shows how a component representing an overall reaction (total_reaction) can encapsulate components representing intermediate reactions (first_reaction and second_reaction) and their by-products (C and D).
<group>
<relationship_ref relationship="is_encapsulated_by" />
<component_ref component="total_reaction" role="major" />
<component_ref component="first_reaction" role="minor" />
<component_ref component="second_reaction" role="minor" />
<component_ref component="C" role="minor" />
<component_ref component="D" role="minor" />
</group>
Figure 11
Example demonstrating the use of the <group> element to define a logical encapsulation relationship. See text for more details.
Figure 12 demonstrates the use of the <group> element to define encapsulation and containment relationships, the construction of two named geometric hierarchies, and the specification of a custom relationship type (is_next_to) in an extension namespace. Most CellML models will probably only define a single geometric hierarchy. In this case, it is not necessary to name the hierarchy, since all unnamed groups are assumed to belong to the same geometric hierarchy.
<group>
<relationship_ref name="membrane" relationship="is_contained_in" />
<component_ref component="cell" role="major" />
<component_ref component="cell_membrane" role="minor" />
</group>
<group>
<relationship_ref relationship="is_encapsulated_by" />
<relationship_ref name="membrane" relationship="is_contained_in" />
<component_ref component="cell_membrane" role="major" />
<component_ref component="sodium_channel" role="minor" />
<component_ref component="calcium_channel" role="minor" />
</group>
<group>
<relationship_ref name="intracellular" relationship="is_contained_in" />
<component_ref component="cell" role="major" />
<component_ref component="network_sarcoplasmic_reticulum" role="minor" />
<component_ref component="junctional_sarcoplasmic_reticulum" role="minor" />
</group>
<group>
<relationship_ref
app:relationship="is_next_to"
xmlns:app="http://www.software.com/cellml_processor" />
<component_ref component="network_sarcoplasmic_reticulum" role="minor" />
<component_ref component="junctional_sarcoplasmic_reticulum" role="minor" />
</group>
Figure 12
Examples demonstrating the use of the <group> element. See text for more details.
The first <group> element states that the cell_membrane component is physically inside the cell component, and that this geometric relationship is part of a geometric hierarchy called membrane. The second <group> element states that the sodium_channel and calcium_channel components are both physically inside and logically encapsulated by the cell_membrane component. This completes the membrane geometric hierarchy. The encapsulation relationship prevents the sodium and calcium channel components from being connected to any components other than the cell_membrane component, each other, and any components they in turn encapsulate.
The third <group> element states that the two components representing parts of the sarcoplasmic reticulum are physically inside the cell, and that this relationship is part of a geometric hierarchy called intracellular. Finally, the fourth <group> element introduces the user-defined relationship is_next_to, and states that the the two sarcoplasmic reticulum components share this relationship. This relationship type is declared by putting the relationship attribute in an extension namespace, and assigning it a value of "is_next_to". Note that this relationship has no major or dominant component, and that CellML processing software is free to ignore the information provided by this group.
- Allowed use of the
<group> element
-
A <model> element may contain any number of <group> elements.
-
A <group> element must contain only the following elements,
which may appear in any order:
-
<relationship_ref> and <component_ref> elements in the CellML namespace,
- metadata framework elements, as described in Section 8.
[ Recommended practice is to define the CellML namespace child elements in a <group> element in the order stated above. ]
-
A <group> element must contain at least one <relationship_ref> element.
-
A <group> element must contain at least one <component_ref> element.
- Allowed use of the
<relationship_ref> element
-
A <relationship_ref> element must contain only the following elements,
which may appear in any order:
- metadata framework elements, as described in Section 8.
-
Each <relationship_ref> element must define a relationship attribute in either the CellML namespace or an extension namespace. It may also define a name attribute.
[ A relationship attribute declaring a user-defined relationship type is placed in an extension namespace. This prevent conflicts with future versions of the CellML specification, which may define additional types of relationships in the CellML namespace. ]
- Allowed values of the
relationship attribute
- Allowed values of the
name attribute
-
The value of the name attribute must be a valid CellML identifier as discussed in Section 2.2.1.
[ Note that unlike most other name attributes, the value of the name attribute on a <relationship_ref> element is not expected to be unique across the current model. Instead, <group> elements that include <relationship_ref> elements that share the same name attribute value form are parts of a single hierarchy. ]
- Proper use of the
name attribute
-
A name attribute may not be defined on a <relationship_ref> element with a relationship attribute value of "is_encapsulated_by".
[ A model may define only a single, unnamed encapsulation hierarchy. ]
- Allowed use of the
<component_ref> element within a <group> element
-
A <component_ref> element must contain only the following elements,
which may appear in any order:
- metadata framework elements, as described in Section 8.
-
A <component_ref> element within a <group> element must define a component attribute and a role attribute.
- Proper use of the
<component_ref> element in <group> elements
-
Two <group> elements that contain <relationship_ref> elements with identical relationship attribute values and undefined name attributes may not reference the same major component.
[ A single level of a hierarchy must only be defined with a single <group> element. It would be much more difficult to assemble a hierarchy from a CellML document if a single level of the hierarchy could be shared among multiple <group> elements. ]
-
Two <group> elements that contain <relationship_ref> elements with identical relationship and name attribute values may not reference the same major component.
[ This rule extends the previous rule to include named hierarchies. ]
-
A component must not be referenced as a minor component more than once in a single grouping hierarchy. All <component_ref> elements with a common component attribute value and a role attribute value of minor must be in different hierarchies.
[ A grouping hierarchy must not be circular. ]
- Allowed values of the
component attribute
-
The value of the component attribute must equal the value of the name attribute of a <component> element contained within the current <model> element.
-
The value of the component attribute on a <component_ref> element must be unique across all <component_ref> elements within the parent <group> element.
[ A component may only appear once within a group. ]
- Allowed values of the
role attribute
- Proper use of the
role attribute
-
A <group> element that contains a <relationship_ref> element with a relationship attribute of "is_encapsulated_by" or "is_contained_in" must contain exactly one <component_ref> element with a role attribute value of "major" and at least one <component_ref> element with a role attribute value of "minor".
[ Groups defining an encapsulation or containment relationship must have exactly one dominant component and at least one minor component. ]
A given model may define multiple geometric containment hierarchies, but may only define one logical encapsulation hierarchy.
A grouping hierarchy is built up from multiple <group> elements based on the value of the name attribute of the <relationship_ref> elements. All <group> elements that contain <relationship_ref> elements that share the same relationship and name attribute values are considered to form a single grouping hierarchy. All <group> elements that contain <relationship_ref> elements that share the same relationship attribute value and do not define name attributes are also considered to form a single grouping hierarchy.
If, after the groups that make up a single hierarchy are assembled, the resulting hierarchy is discontinuous, it may be convenient to assume that any components that are not already children of other components are children of a single imaginary component. The imaginary component has no properties in the model. Its sole purpose is to make it easier to check that the hierarchy has no circular relationships between components.
Modellers are explicitly forbidden from using CellML groups to add mathematical information to the model. Modellers may not define their own types of relationships that imply mathematics.
Modellers should not use CellML groups to associate properties or classification information with sets of components. The metadata functionality is the proper method for making such associations. This increases the chance of that information being used by a range of CellML processing software.
|