|  |  |  |  |
|  |  |  |  |  | |  |  |  |  |  | | Final Draft — 18 May 2001 | |
Any model can be described as a network of connections between self-contained components. A component is a functional unit that may correspond to a physical compartment, event, or species or may be just a convenient modelling abstraction. A component contains variables and mathematical relationships that manipulate those variables. Connections exchange information between components. A connection contains mappings between variables in two components, allowing the value of a variable in one component to be passed to a variable in the other component.
A model is declared in CellML with a <model> element. This is the usual root element for a CellML document. Throughout this section of the specification, all elements are in the CellML namespace unless stated otherwise. The recommended method for specifying namespaces in a CellML document is described in Section 2.2.3.
The <model> element has a name attribute that allows this model to be unambiguously referenced by other models. For instance, this would be necessary if the model were to be combined with other models or partial models to create a larger model.
A <model> element may contain any number of the elements in the following list in any order. The recommended practice is for elements placed within the <model> element to appear in the order given in the following list. This allows people to quickly find certain kinds of information within a CellML document.
-
<units> — A modeller can declare a set of units to use in the model, as described in Section 5.
-
<component> — Components are the smallest functional units in a model. Each component contains variables that represent the key properties of the component and mathematics that describe the behaviour of the portion of the system represented by that component.
-
<group> — Groups allow the modeller to define logical and physical relationships between components. Groups are defined using the <group> element, as discussed in Section 6.
-
<connection> — Connections are used to connect components to each other and to map variables in one component to variables in another. Connections are defined using the <connection> element, as discussed in Section 3.2.4.
The <model> element (and indeed any of the elements in a CellML document) may define metadata to provide context for that object. This metadata might include documentation, citations from literature, or a modification history for the current CellML object. Adding metadata to a CellML document is discussed in detail in Section 8.
Constructing a model from multiple components encourages the re-use of components. For instance, an electro-physiological model of a cell might be organised into components that represent various ion channels. All of the mathematics that describe the behaviour of the L-type calcium channel would be defined in a single component representing this particular ion channel. If a modeller wished to re-use the portion of the model representing the L-type calcium channel in another model, he or she would only need to copy this component.
A <component> element is used to declare a CellML component. It must only be used inside a <model> element or as the root element of a CellML document. A <component> element that is the root of a CellML document does not define a complete model. It would probably be part of a library of standard components that could be imported and used in models. Eventually, CellML will include a mechanism that simplifies such re-use of components. At the present time, the component would need to be physically copied into a model document to be used in that model.
A CellML <model> may contain any number of <component> elements. Each <component> must have a name attribute, the value of which is a unique identifier for the component within the current model. The value of the name attribute is used to reference the component in other parts of the model, such as in connections and groups.
A <component> may contain any of the elements in the following list in any order. Again, recommended practice is for elements placed within the <component> element to appear in the order given in the following list.
-
<units> — A modeller can declare a set of units to use within the component, as described in Section 5.
-
<variable> — A component may contain any number of <variable> elements, which define variables that may be mathematically related in the equation blocks contained in the component. Variables are discussed in Section 3.2.3.
-
<reaction> — A component may contain <reaction> elements, which are used to provide chemical and biochemical context for the equations describing a reaction. It is recommended that only one <reaction> element appear in any <component> element. The definition of reaction information is described in Section 7.
-
<mathml:math> — A component may contain a set of mathematical relationships between the variables declared in this component. These equations are marked up using MathML, as discussed in Section 4. The mathml prefix is used to indicate that the <math> element is in the MathML namespace.
A <component> element is also a sensible place to define metadata, using the syntax presented in Section 8.
The definitions of two <component> elements are included in the example described in Section 3.3.
Models are usually developed to simulate the behaviour of a number of variables that have physiological significance. Each variable in the model belongs to a single component, which may contain equations that modify the value of that variable. The value of a variable may be passed through connections into other components. The variable must also be declared in these components, which can then use the value of the variable in their own equations but must not modify it.
The <variable> element is used to declare a CellML variable. It can only be used inside a <component> element. Variables must define a name attribute, the value of which must be unique across all variables in the current component. The name of a variable is used when referencing variables inside connections (see Section 3.2.4) and reactions (see Section 7). All variables must also define a units attribute. The value of this attribute must correspond to one of the keywords in the CellML units dictionary or the value of the name attribute of a units element defined within the current component or model, as described in Section 5.
A <variable> element may also have the following attributes:
-
initial_value — This attribute provides a convenient means for specifying the initial or default value of a scalar variable in a simulation with time as the independent variable. The variable's value may be reset or modified in equations in the current component. The initial values of variables need not be set in the model definition at all; they could instead be set in a configuration file loaded separately by the model processor.
-
public_interface — This attribute specifies the interface exposed to components in the parent and sibling sets (see below). The public interface must have a value "in", "out", or "none". The absence of a public_interface attribute implies a value of "none".
-
private_interface — This attribute specifies the interface exposed to components in the encapsulated set (see below). The private interface must have a value "in", "out", or "none". The absence of a private_interface attribute implies a value of "none".
Whether or not a component may obtain the value of a variable in another component depends on the public_interface and private_interface attributes on the variable declaration, and the place of the two components in the encapsulation hierarchy.
Encapsulation allows the modeller to hide a complex network of components from the rest of the model and provide a single component as an interface to the hidden network. Encapsulation effectively divides the network into layers, where connections between the layers must only be made through the interface components.
The components to which any given component may connect can be divided into four distinct sets with respect to any given component (the current component). The set of all components immediately encapsulated by the current component is referred to as the encapsulated set. If the current component is encapsulated, then the encapsulating component is referred to as the parent, and the set of all other components encapsulated by the same parent is referred to as the sibling set. If the current component is not encapsulated, then it has no parent and the sibling set consists of all other components in the model that are not encapsulated. All other components, which are not available to make connections with the current component, make up the hidden set. The encapsulation hierarchy and its effects on variable mapping are described in Section 6.
When a variable is declared with either a public_interface or private_interface attribute value of "in", then the value of that variable must be imported from another component. Otherwise, a variable's value must be set and modified in the current component. The variable is then said to belong to or be owned by the current component.
Eventually, it will be possible to specify the temporal and/or spatial variation of a variable's value using FieldML. The capability to include FieldML is still under development. At the present time, all variables must have singular values.
Connections provide the mechanism for mapping variables declared within one component to variables in another component, allowing information to be exchanged between the various components in the network. The mapping of variables involves the transfer of a variable's value from one component to another. There will be many such mappings present in a network. The transfer of a variable's value may involve a conversion to account for the units in which each component expects the variable's value to be defined. (More information on units conversion can be found in Section 5.)
The complete set of variable mappings between any two components constitutes a connection. Only one connection may be created between any given pair of components in a model. Each connection references the two components involved in the connection, and then maps variables from each of the components together. It is not necessary for the variables that are to be mapped to each other to have the same name. However, the interface attributes of each pair of variables must be compatible — an "out" variable in one component's interface must map to an "in" variable in the other component's interface. The direction of each mapping is determined by the value of the public_interface and private_interface attributes on the two variables: the value is always passed from the variable with an interface value of "out" to the variable with an interface value of "in". The value of a variable declared with an interface value of "out" may be passed out to any number of variables in other components declared with interface values of "in". The component to which a variable belongs is found by following the variable back from "in" to "out" interfaces, following the model's connections.
The <connection> element is used to declare a CellML connection. It can only appear inside a <model> element.
A <connection> element must contain exactly one <map_components> element, which is used to reference the two components involved in the connection. Each <map_components> element must define component_1 and component_2 attributes, the values of which are the names of the components being referenced. In CellML 1.0, the referenced components must be defined within the current <model> element. It is anticipated that it will eventually be possible to reference components from other models, allowing models to be connected into larger models.
A <connection> element must also contain one or more <map_variables> elements, which are used to reference the variables being mapped between the two components in the connection. Each <map_variables> element must define variable_1 and variable_2 attributes, the values of which are equal to the names of variables defined in the components referenced by the component_1 and component_2 attributes on the <map_components> element, respectively.
The CellML example discussed in Section 3.3 demonstrates the definition of a <connection> element.
Figure 4 contains a portion of the CellML encoding of the Hodgkin-Huxley squid axon model published in 1952. The excerpt contains the definitions of the components corresponding to the membrane and the sodium channel, and the connection between the two components. Most of the complexity from the full model definition has been left out for conciseness and clarity. This example is only used to demonstrate the standard use of the <component>, <variable>, and <connection> elements.
<model
name="hodgkin_huxley_model_excerpt"
xmlns="http://www.cellml.org/cellml/1.0#"
xmlns:cellml="http://www.cellml.org/cellml/1.0#"
xmlns:cmeta="http://www.cellml.org/metadata/1.0#">
<component name="membrane">
<!-- the following variable is used in other components -->
<variable
name="V" initial_value="-75.0"
public_interface="out" units="millivolt" />
<!-- the following variables are imported from other components -->
<variable name="time" public_interface="in" units="millisecond" />
<variable name="i_Na" public_interface="in" units="microA_per_cm2" />
<variable name="i_K" public_interface="in" units="microA_per_cm2" />
<variable name="i_L" public_interface="in" units="microA_per_cm2" />
<!-- the following variable is only used internally -->
<variable name="C" initial_value="1.0" units="microF_per_cm2" />
<math xmlns="http://www.w3.org/1998/Math/MathML">
<apply id="voltage_current_relationship"><eq />
<apply><diff />
<bvar><ci> time </ci></bvar>
<ci> V </ci>
</apply>
<apply><divide />
<apply><plus />
<ci> i_Na </ci>
<ci> i_K </ci>
<ci> i_L </ci>
</apply>
<ci> C </ci>
</apply>
</apply>
</math>
</component>
<component name="sodium_channel">
<!-- the following variables are used in other components -->
<variable name="i_Na" public_interface="out" units="microA_per_cm2" />
<!-- the following variables are imported from other components -->
<variable name="time" public_interface="in" units="millisecond" />
<variable name="V" public_interface="in" units="millivolt" />
<math xmlns="http://www.w3.org/1998/Math/MathML">
<apply id="i_Na_calculation"><eq />
<ci> i_Na </ci>
... <!-- a function of V & time -->
</apply>
</math>
</component>
<connection>
<map_components component_1="membrane" component_2="sodium_channel" />
<map_variables variable_1="V" variable_2="V" />
<map_variables variable_1="i_Na" variable_2="i_Na" />
</connection>
</model>
Figure 4
A small portion of the CellML definition of the Hodgkin-Huxley squid axon model from 1952. This excerpt contains the definition of the components corresponding to the membrane and the sodium channel, and the connection between them. Much detail has been omitted, but this example clearly demonstrates the relationship between the <component>, <variable> and <connection> elements. See the text for more information.
The membrane component declares six variables, which are divided into three categories. The first variable is called V, and it represents the membrane voltage in the model. It has a public_interface attribute value of "out", which indicates that the variable "belongs" to this component and that its value may be obtained by other components in the model via connections. It references a units definition by the name of millivolt (this definition is not included here), and is given an initial value of -75.0 millivolts.
The subsequent four variables are time, i_Na (sodium current), i_K (potassium current) and i_L (leakage current). They are all declared with a public_interface attribute value of "in", which indicates that their values are obtained from other components via connections.
Finally, a variable C (capacitance) is declared. This <variable> element defines neither a public_interface or a private_interface attribute. Both of these attributes therefore assume the default value of "none", which means that the variable belongs to the current component and is not visible to other components in the model.
After the variable declarations, a <math> element in the MathML namespace is used to define an equation relating V to the other variables. Only the values of the variables belonging to a component may be mathematically modified in that component. The full set of equations defined in the membrane component is too lengthy to include here. The equation included in Figure 4 is the well known differential equation from the Hodgkin Huxley model:
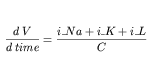 | (1) |
The sodium_channel component declares three variables, all of which represent quantities that were also declared in the membrane component. The i_Na variable declared in this component has a public_interface attribute value of "out", indicating that the sodium current belongs to this component. The value of the sodium current is calculated in this component, although the actual math has been omitted.
Finally, a <connection> element references the membrane and sodium_channel components using a <map_components> element, and maps the V and i_Na variables in each component together, using two <map_variables> elements. The value of the variable_1 attribute on each <map_variables> element references the corresponding variable in the membrane component, which is the component referenced by the component_1 attribute on the <map_components> element. Similarly, the values of the variable_2 attributes reference variables in the sodium_channel component.
The following are the rules for using the <model>, <component>, <variable>, <connection>, and <map_components> elements, and <map_variables> elements.
- Allowed use of the
<model> element
-
A <model> element must contain only the following elements,
which may appear in any order:
-
<units>, <component>, <connection>, and <group> elements in the CellML namespace,
- metadata framework elements, as described in Section 8.
[ Recommended practice is to define the child elements in the CellML namespace in the order stated above. ]
-
Each <model> element must define a name attribute.
- Allowed values of the
name attribute
- Allowed use of the
<component> element
-
A <component> element must contain only the following elements,
which may appear in any order:
<units> and <variable> elements in the CellML namespace,
-
<math> elements in the MathML namespace,
- metadata framework elements, as described in Section 8.
[ Recommended practice is to define the child elements in the CellML and MathML namespaces in the order stated above. Note that a <component> element must not appear inside another <component> element. Such nesting could be intended to indicate a logical encapsulation relationship, a geometric containment relationship, or some other relationship between the two components. There is no reason to assume that the nesting hierarchy produced for one type of relationship would be consistent with the hierarchy produced for other types of relationship. Therefore, CellML defines these relationships using the <group> element, rather than nesting of <component> elements. ]
-
Each <component> element must define a name attribute.
- Allowed values of the
name attribute
-
The value of the name attribute must be a valid CellML identifier as discussed in Section 2.2.1.
-
The value of the name attribute must be unique across all <component> elements contained in the parent <model> element.
- Allowed use of the
<variable> element
-
A <variable> element must contain only the following elements,
which may appear in any order:
- metadata framework elements, as described in Section 8.
-
Each <variable> element must define a name attribute and a units attribute. It may also define public_interface, private_interface, and initial_value attributes.
- Allowed values of the
name attribute
-
The value of the name attribute must be a valid CellML identifier as discussed in Section 2.2.1.
-
The value of the name attribute of a <variable> element must be unique across all <variable> elements contained in the same <component> element.
[ Two variables in the same component must not have the same name. However, two variables in different components may have the same name, and a variable may have the same name as its parent component. ]
- Allowed values of the
units attribute
-
The value of the units attribute must either be one of the keywords defined in the standard dictionary or the value of the name attribute on a <units> element defined in the current component or model.
[ The dictionary and the units element are described in Section 5. ]
- Allowed values of the
public_interface attribute
-
If present, the value of the public_interface attribute must be "in", "out", or "none".
-
If not present, its value defaults to "none".
- Allowed values of the
private_interface attribute
-
If present, the value of the private_interface attribute must be "in", "out", or "none".
-
If not present, its value defaults to "none".
- Proper use of the
public_interface and private_interface attributes
- Allowed values of the
initial_value attribute
-
If present, the value of the initial_value attribute must be a real number.
-
The absence of an initial_value attribute implies nothing.
[ The absence of this attribute would usually mean either that the variable does not need an initial value or that this value will be supplied in a parameter file or by the user at the time simulations using the model are run. ]
- Proper use of the
initial_value attribute
-
An initial_value attribute must not be defined on a <variable> element with a public_interface or private_interface attribute with a value of "in".
[ These variables receive their value from variables belonging to another component. ]
- Allowed use of the
<connection> element
-
A <connection> element must contain only the following elements,
which may appear in any order:
<map_components> and <map_variables> elements in the CellML namespace,- metadata framework elements, as described in Section 8.
-
Each <connection> element must contain exactly one <map_components> element.
-
Each <connection> element must contain at least one <map_variables> element.
[ It does not make sense to define a connection that does not map variables together. This rule prevents software from using empty connections to imply information not defined in this specification. ]
- Allowed use of the
<map_components> element
-
A <map_components> element must contain only the following elements,
which may appear in any order:
- metadata framework elements, as described in Section 8.
-
Each <map_components> element must define a component_1 attribute and a component_2 attribute.
- Allowed values of the
component_1 attribute
- Allowed values of the
component_2 attribute
- Proper use of the
component_1 and component_2 attributes
-
The component_1 and component_2 attributes on a single <map_components> element must not have the same value.
[ A connection must link two different components. ]
-
Each <map_components> element contained within <connection> elements that are contained within a given <model> element must define a unique pair of component_1 and component_2 attribute values.
[ There can only be one connection between any two components in a network. This prevents setting up inconsistent, circular, or duplicate variable mappings between any two components in the network. However, it does not prevent a model author from creating inconsistent mathematical relationships between the variables. ]
- Allowed use of the
<map_variables> element
-
A <map_variables> element must contain only the following elements,
which may appear in any order:
- metadata framework elements, as described in Section 8.
-
Each <map_variables> element must define a variable_1 attribute and a variable_2 attribute.
- Allowed values of the
variable_1 attribute
- Allowed values of the
variable_2 attribute
- Proper use of the
<map_variables> element to map variables to each other
[ The rules for mapping a variable to other variables depend on the encapsulation hierarchy of the component that owns the variable. This hierarchy divides the rest of the components in the model into parent, sibling, encapsulated, and hidden sets, as described in Section 3.2.3. The public_interface attribute defines the availability of a variable to the parent component and components in the sibling set. The private_interface attribute defines the availability of a variable to components in the encapsulated set. Variables are not available to components in the hidden set. ] -
Variables with a public_interface or private_interface attribute value of "in" must be mapped to variables with a public_interface or private_interface attribute value of "out".
-
A variable with either a private_interface or public_interface attribute value of "in" must be mapped to no more than one other variable in the model.
[ Note that a similar restriction does not apply to variables with interface values of "out". An output variable can be mapped to multiple input variables in various components in the current model. ]
-
A variable with a public_interface attribute value of "in" must be mapped to a single variable owned by a component in the sibling set, provided the target variable has a public_interface attribute value of "out", or to a single variable owned by the parent component, provided the target variable has a private_interface attribute value of "out".
-
A variable with a public_interface attribute value of "out" may be mapped to variables owned by components in the sibling set, provided the target variables have public_interface attribute values of "in". It may also be mapped to variables owned by the parent component, provided the target variables have private_interface attribute values of "in".
-
A variable with a private_interface attribute value of "in" may be mapped to a single variable owned by a component in the encapsulated set, provided the target variable has a public_interface attribute value of "out".
-
A variable with a private_interface attribute value of "out" may be mapped to variables owned by components in the encapsulated set, provided the target variables have public_interface attribute values of "in".
-
A
<map_variables> element maps together two variables from the components referenced by the component_1 and component_2 attributes on the <map_components> element within the current <connection> element. The variable_1 attribute references a variable defined in the component referenced by the component_1 attribute, and the variable_2 attribute references a variable defined in the component referenced by the component_1 attribute. Variable mappings specifically do not depend on variable names.
|  |  |  |  |  |