CellML Graph Metadata Specification - 02
- PLEASE NOTE: development of this specification is currently on-hold in favour of developing SED-ML to meet the requirements originally being addressed via the approach specified below.
- This version:
- http://www.cellml.org/specifications/metadata/graphs/draft-graph-metadata-02
- Latest version:
- http://www.cellml.org/specifications/metadata/graphs
Authors:
- David Nickerson <david.nickerson@nus.edu.sg> (Division of Bioengineering, National University of Singapore)
- Andrew Miller <ak.miller@auckland.ac.nz> (Auckland Bioengineering Institute, The University of Auckland)
Abstract
CellML provides a mechanism to describe mathematical models. While CellML can describe the mathematics used by many models published in scientific journals, it does not describe all of the information required to produce graphical outputs illustrating the behaviour of those models. One step towards this is reproducing experimental results, and this is described in the simulation metadata specification. However, it is also useful to be able to produce the exact graphs from a paper, and this specification allows for metadata which provides the information needed to do this.
Such data fits naturally into a CellML model, as metadata. CellML metadata is expressed in Resource Description Format (RDF). This specification provides an RDF based language for describing graphs. As RDF is an open-world framework, such RDF data may be embedded in a CellML document, or alternatively, could be provided by a third party in a separate document.
Status of this document
This document is a discussion draft version of the CellML graph metadata specification, and has not been endorsed by the CellML team or members of the CellML community. Comments are solicited and should be sent to cellml-discussion@cellml.org.
Introduction
CellML defines a mathematical model and simulation metadata defines a specific instantiation of that model in terms of a numerical simulation run. The graphing metadata we define here specifies a method to combine simulation results from one or more simulations performed using one or more CellML models to provide a graphical representation of the model behaviour in the form of two-dimensional graphs. The most obvious use of such metadata is to unambiguously define machine readable descriptions of graphs for use in publications, but they may also prove useful in some user interface applications.
There are three main aspects to this graphing metadata specification:
- extracting data sets from simulation results;
- the graphical representation of a data set; and
- the combination of graphical data representations into a (possibly annotated) graph.
The first of these are defined herein as graph variables which are encapsulated by traces. Traces define the data for the graphical representation of a pair of graph variables. Collections of traces combine to make a graph which accomplishes the third aspect above.
An additional aspect to those above is the association of experimental data to simulation results. This fits naturally within trace definitions as demonstrated below.
Graphical summary of core features (non-normative)
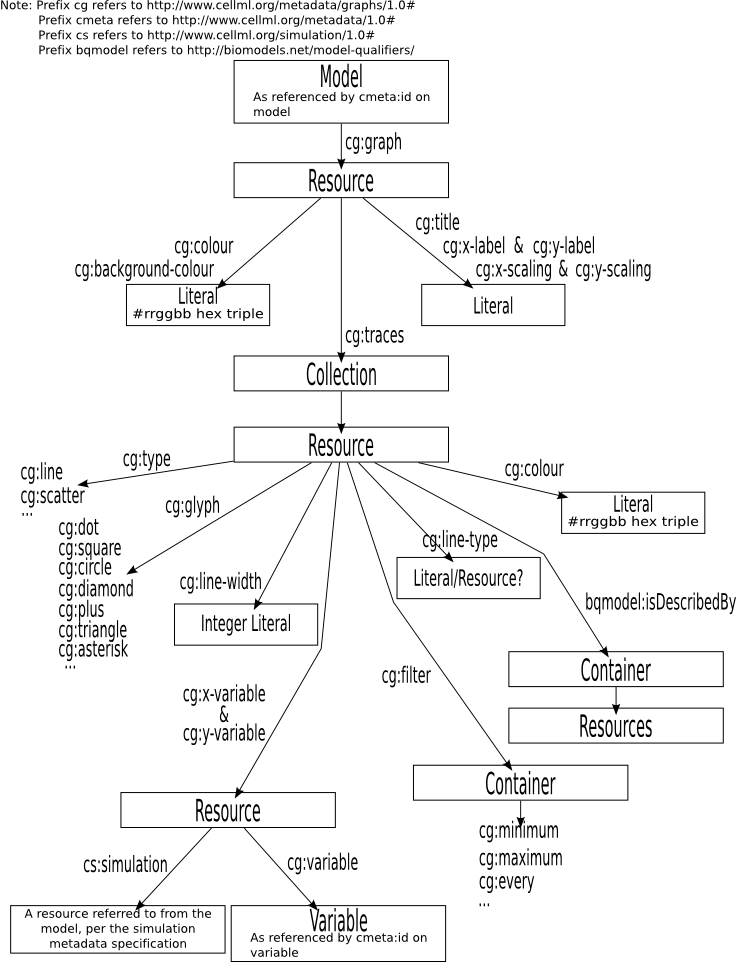
Prefix
The URI http://www.cellml.org/metadata/graphs/1.0# is used as the prefix for all RDF predicates defined in this specification. When describing documents in RDF/XML, it is recommended that the prefix cg be bound to this URI. However, processing software which parses CellML graph metadata in RDF/XML form must not assume that this prefix will be used.
Within this document, predicate resources are referred to using the notation of qualified names in RDF/XML, and assuming that the cg prefix is bound to "http://www.cellml.org/metadata/graphs/1.0#". For example, cg:graph refers to the resource "http://www.cellml.org/metadata/graphs/1.0#graph".
Within this document, it is assumed that the prefix cmeta is bound to the XML namespace "http://www.cellml.org/metadata/1.0#", that the prefix cs is bound to the XML namespace "http://www.cellml.org/metadata/simulation/1.0#", and that the prefix bqmodel is bound to the XML namespace "http://biomodels.net/model-qualifiers/". Processing software, however, must not assume that these prefixes will always be used.
Specifications of graphs
As described in the CellML metadata specification, the CellML model is associated with the RDF resource with fragment identifier matching the cmeta:id on the model element and the URI base equal to the CellML document URI.
Every CellML model may have zero or more graphs associated with them. Such graphs are specified by an arc from the model (subject) to a node (object), with predicate cg:graph. As one model can have more than one graph associated with it, it is not a contradiction for two or more such arcs to exist. Where CellML processing software discovers more than one such arc, it may display them all, or may prompt the user to select one or more, or it may choose one or more arcs to display arbitrarily.
Throughout this document, the object node of the arc just described is referred to as the "graph node".
Graphs which refer to more than one model may also be defined. In this case, the mechanism by which processing software discovers the graph node is beyond the scope of this specification. In this case it is recommended that the graph node is described externally to all the CellML models it refers to.
The graph node (and all other nodes reachable from it by arcs defined in this specification) shall be closed-world with respect to predicates defined in this specification — that is, it shall be completely specified within the XML file in which it is defined. If a user wishes to add additional details defined in this specification to a different XML document, they must not attempt to define new arcs with the graph node (or any other nodes reachable from it defined in this specification) as the subject. Instead, they should define a new graph with the additional information. However, it is acceptable for predicates which annotate the graph(perhaps, for example, to comment on the relationship between the graph and reality), rather than define it, to be specified in a different XML file.
A graph is made up of one or more traces superimposed on top of each other on a set of axes. A common numerical scale on each axis is shared by all traces (even if the units are incompatible). The implementing software is free to choose this numerical scale however it likes. Future versions of this specification may allow the metadata to specify a scale for the software to use. [it is quite common to define two independent y-axes on a single graph (e.g., current on one and concentration on the other). This is typically achieved through the introduction of a "y2" axis. Such a concept should be fairly easy to insert into this specification if we think it is worth having.]
A graph node may also optionally define the predicates:
- cg:title
- A human readable literal string which processing software may use to define a title for the graph.
- cg:background-colour
- A string literal consisting of a standard hexadecimal triple (#RRGGBB) specifying a preferred background colour for the graph. Processing software supporting the cg:background-colour and cg:colour properties is expected to select reasonable default values if only one or other of these properties is defined for a graph. [perhaps we should also allow the standard HTML colour string names for all current hex triples?]
- cg:colour
- A string literal consisting of a standard hexadecimal triple (#RRGGBB) specifying the preferred foreground colour for the graph. If processing software supports this property it is expected to also be used for all traces not specifying their own forground colour. Processing software supporting the cg:colour and cg:background-colour properties is expected to select reasonable default values if only one or other of these properties is defined for a graph.
- cg:x-label & cg:y-label
- Human readable literal strings which processing software may use to label the x and y axes.
- cg:x-scaling & cg:y-scaling
- String literals defining scaling to be applied to either the x or y values of the graph. Processing software supporting this property is expected to clearly define the types of scaling supported. [not too sure about this one, it makes sense to have it here but we need to probably define the strings for different types of scaling that can be used. so far have only considered the case of log10 scaling.]
When describing graph nodes in RDF/XML, it is recommended that the node be given an explicit resource URL, rather than using the anonymous node facilities. This makes it easier for other RDF documents to refer to the graph.
Specification of traces
For each graph node, there shall be exactly one arc with subject equal to the graph node, and predicate equal to cg:traces. The object of this node shall be a collection. Each member of the collection shall be a resource. Throughout this document, the members of the collection just described are referred to as the "trace nodes".
Each trace node shall have exactly one arc with subject equal to the trace node, and predicate equal to cg:type. The object of this arc shall be a resource taken from a limited set. This specification defines two resources, cg:line (for a line graph) and cg:scatter (for a scatter plot), which may be used as the object of this arc. Other specifications may add additional resources.
Each trace node of type cg:line or cg:scatter shall have exactly one arc with subject equal to the trace node, and predicate cg:x-variable, and similarly for the predicate cg:y-variable. The objects of these arcs shall be resources. Such resources shall be referred to as trace variable nodes within this specification.
Each trace node shall have zero or one arc with subject equal to the trace node and predicate equal to cg:label. The object of this arc shall be a plain literal. This property specifies a human readable label for the trace which processing software may use in labelling data within a graph (a graph key or similar, for example). [perhaps cg:title is more appropriate?]
Each trace node shall have zero or one arc with subject equal to the trace node, and predicate equal to cg:colour. The object of this arc shall be a plain literal. The literal shall be formatted as a # character, followed by six hex digits (made up of the character 0-9, A-F, and a-f). The first two hex digits shall specify the red component of the colour of the trace, with the second two specifying the green component, and the final two specifying the blue component. If supported, processing software is expected to use this colour when drawing the trace data. If not specified for the trace node, processing software is expected to fall back on the cg:colour property of the parent graph node. If no cg:colour property is found processing software is free to determine a suitable colour for the trace data.
Each trace node of type cg:scatter shall have zero or one arc with subject equal to the trace node, and predicate equal to cg:glyph. The object of this arc shall be a resource taken from a limited set. This specification defines seven glyphs, cg:dot, cg:square, cg:circle, cg:diamond, cg:plus, cg:triangle, and cg:asterisk. Other specifications may add additional resources. If not present processing software is free to choose a suitable glyph graphic for use in drawing scatter plots. [As with cg:line-type, cg:glyph could just be an integer to indicate to processing software what should have the same glyph and what should be different. The problem with this approach is that it is often necessary to indicate specific data from within article text so its good to know what (fixed) symbol will be used.]
Each trace node of type cg:line shall have zero or one arc with subject equal to the trace node and predicate equal to cg:line-type. The object of this arc shall be an integer literal. Processing software supporting this property is expected to use a different line type (solid, dashed, dotted, etc.) for each different cg:line-type value used within a single graph node. Identical cg:line-type values are expected to give the same graphical line drawing within a single graph. Similarly, all trace nodes of type cg:line which do not specify a cg:line-type are expected to be drawn with the same graphical line type. [maybe rather than an integer literal this should be a resource and we define the standard line types that applications can use? cg:solid, cg:dashed, cg:dotted, cg:dash-dot, etc.]
Each trace node of type cg:line shall have zero or one arc with subject equal to the trace node and predicate equal to cg:line-width. The object of this arc shall be an integer literal. Processing software supporting this property is expected to draw increasing thickness lines for increasing values of cg:line-thickness.
Each trace node shall have zero or one arc with subject equal to the trace node and predicate cg:filter. The object of this arc shall be a container. Each member of the container shall be a filter to be applied to the trace data. The type of container used indicates to supporting software the order in which the filters are applied (i.e., if order is important a rdf:Seq type container should be used). This specification defines the filters: cg:minimum, cg:maximum, cg:sort, and cg:every. Some of these filters need to refer to the trace variable node to be used in the filtering process....need to define the filters and expected behaviour from processing software. [cg:minimum and cg:maximum are required to pull out graphs of a subset of data - possibly only makes sense to do this based on the x-variable, but we should let processing software decide that. I haven't found a use for cg:sort yet. cg:every is useful to pull out an evenly spaced subset of data spanning the entire range - i.e., if you have a simulation of 75 minutes worth with data every millisecond you get a huge graph if you draw the full data set.]
Each trace node shall have zero or one arc with subject equal to the trace node and predicate equal to bqmodel:isDescribedBy. The object of this arc shall be a container (rdf:Bag, rdf:Seq, or rdf:Alt). Each member of the container shall be a resource and each resource shall be a reference to data external to this specification which can be used by processing software to apply additional information to this trace. Experimental data or previous simulation results are two such sources of data that are thought to be useful here. [this definitely needs to be stated better and more clearly]
Specification of trace variables
Each trace variable node shall have exactly one arc with subject equal to the trace variable node, and predicate cg:variable. The object of this arc shall be an RDF resource with fragment identifier matching the cmeta:id on a variable element and the URI base equal to the CellML document URI in which the variable is found.
Each trace variable node shall have exactly one arc with subject equal to the trace variable node, and predicate cs:simulation. The object of this arc shall be a simulation node resource, as defined in the simulation metadata specification.
When describing trace nodes in RDF/XML, it is recommended that the node be given an explicit resource URL, rather than using the anonymous node facilities. This makes it easier for other RDF documents to refer to the trace.
Appendix A: Sample graph metadata (non-normative)
See the examples for some sample graph metadata.

